Here, we demonstrate that with the increased sensitivity of modern FTIR spectrometers, automation (check this) of sample preparation and modern bioinformatics, it is possible to identify and validate spectral biomarker candidates so that we can distinguish between urinary bladder cancer (UBC) and inflammation in suspected bladder cancer patients.
The current dataset contains spectra of blood serum and plasma samples of 135 patients. All patients underwent cytology and pathological biopsy characterisation to distinguish between patients without UBC (46) and confirmed UBC cases (89). A minimally invasive blood test could spare control patients a repeated cystoscopy, which includes a transurethral biopsy and a three-day stationary hospitalisation.
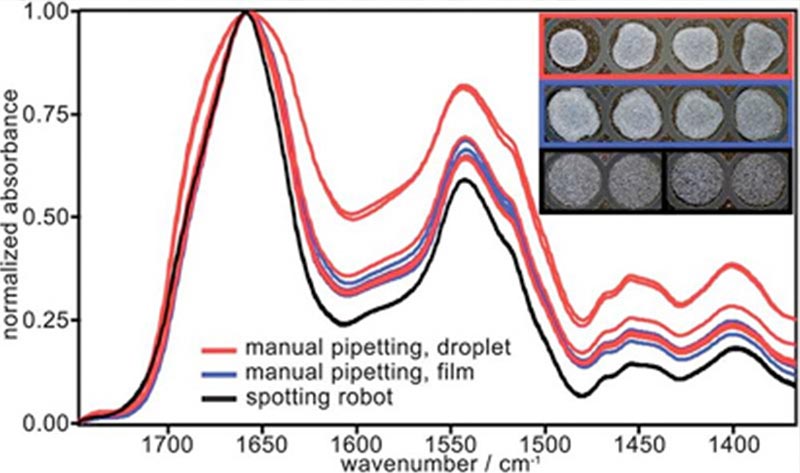
Blood serum, EDTA and citrate plasma were collected from each patient and processed following predefined strict standard operating procedures. Highly reproducible dry films were obtained by spotting subnanolitre biofluid droplets in defined patterns, which were then compared and optimised. Particular attention was paid to the automation of sample preparation and spectral pre-processing to exclude errors by manual handling. Spectral biomarker candidates were identified from absorbance spectra, as well as their first and second derivative spectra, using an advanced Random Forest (RF) approach.
It turned out that the second derivative spectra were most useful for classification. Repeat validation on 21% of the dataset not included in predictor training with Linear Discriminant Analysis (LDA) classifiers and Random Forests (RFs) yielded a sensitivity of 93 _ 10% and a specificity of 46 _ 18% for bladder cancer.
The low specificity can be most likely attributed to the unbalanced and small number of control samples. Using this approach, spectral biomarker candidates in blood-derived biofluids were identified, which allow us to distinguish between cancer and inflammation, but the observed differences were tiny. Obviously, a much larger sample number has to be investigated to reliably validate such candidates.
Author: Julian Ollesch, Steffen L. Drees, H. Michael Heise, Thomas Behrens,Thomas Brüning and Klaus Gerwert